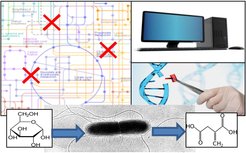
Metabolic Engineering of microbial cell factories
We combine computational (dry-lab; see research area 1) and genetic (wet-lab; see research area 2) techniques to develop and implement new metabolic engineering strategies and to build microbial cell factories for the production of selected chemicals. We focus on E. coli as production organism, but are also interested in applications with other relevant microbial hosts such as Zymomonas mobilis. Research topics include:
- Enforced ATP wasting as a metabolic engineering strategy to enhance the productivity of microbial producer strains (see ERC Consolidator project StrainBooster).
- Model-based metabolic engineering of E. coli for synthesis of bulk chemicals (such as itaconic acid, succinate, 2,3-butanediol, isobutanol, isopropanol and others).
- Metabolic redesign of Zymomonas mobilis for efficient synthesis of chemicals beyond ethanol.
- Metabolic engineering strategies for cyanobacteria.
- Use of alternative substrates (e.g. glycerol and acetate) and feedstocks (e.g. biogenic residues such as molasses) beyond sugars.
- Design of cell factories for two-stage processes via dynamic metabolic engineering (see also research area 4).
