
Method Development for Glycoproteomic Analysis
For site-specific glycosylation analyses, glycopeptide-based glycoproteomics using mass spectrometry is the method of choice. The aim of such analyses is the identification of occupied glycosylation sites, including the quantification of the site occupancy (macroheterogeneity), as well as identification, structural characterization and quantification of all glycans attached to the individual glycosylation sites (microheterogeneity). Over the last years we have developed and established mass-spectrometry-based methods and workflows that enable the site-specific and in-depth analysis of N- and O-glycopeptides.
Improvement of the glycoproteomic toolbox with Flavastacin
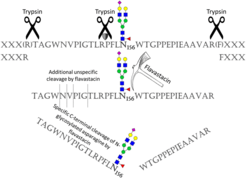
Figure 1: Proteolytic cleavage specificity of trypsin combined with Flavastacin, which cleaves at the C-terminus of N-glycosylated asparagine (Taken from Pralow et al. (2018), Sci Rep).
Among others, progress has been made in the use of different proteolytic enzymes to generate (glyco)peptides. Flavastacin is a protease that is known to cleave N-terminal to aspartic acid residues. However, for the first time, our group discovered a new and unique C-terminal cleavage specificity of Flavastacin for N-glycosylated asparagine (Figure 1). Implemented in an N-glycoproteomic workflow the use of Flavastacin can thus not only render data analysis much easier, it can also significantly increase the confidence of MS-based N-glycoproteomic analyses. Currently we are working on a more profound understanding of the mechanism and specificity of Flavastacin.
LC-MS-based in-depth analysis of glycopeptides
![Figure 2: Top Left: Glycoproteomic workflow developed for the in-depth analysis of N- and O-glycosylated proteins. Top Right: Conserved N-glycopeptide fragmentation pattern consistently obtained when using stepped normalized collisional energy (HCD.step). Bottom: HCD fragment ion spectra (MS2) of tryptic IgG 1 Fc N-glycopeptide 176EEQYNSTYR184 + HexNAc4Hex3Fuc1 (m/z 878.687 [M+3H]3+) acquired by nano-RP-LC-ESI-OT-OT-MS/MS using varying normalized collisional energies (NCE; positive ion mode, LTQ Orbitrap Elite hybrid mass spectrometer). Taken from Hoffmann et al. (2018), Proteomics, in press.](/3629800/original-1547802835.jpg?t=eyJ3aWR0aCI6MjQ2LCJvYmpfaWQiOjM2Mjk4MDB9--c517643091a104d4e25ad81d6699ff80411ac2e3)
Figure 2:
Top Left: Glycoproteomic workflow developed for the in-depth analysis of N- and O-glycosylated proteins. Top Right: Conserved N-glycopeptide fragmentation pattern consistently obtained when using stepped normalized collisional energy (HCD.step). Bottom: HCD fragment ion spectra (MS2) of tryptic IgG 1 Fc N-glycopeptide 176EEQYNSTYR184 + HexNAc4Hex3Fuc1 (m/z 878.687 [M+3H]3+) acquired by nano-RP-LC-ESI-OT-OT-MS/MS using varying normalized collisional energies (NCE; positive ion mode, LTQ Orbitrap Elite hybrid mass spectrometer). Taken from Hoffmann et al. (2018), Proteomics, in press.
We have developed a glycoproteomic workflow that enables the site-specific and in-depth glycosylation analysis of both N- and O-glycopeptides (Figure 2). The workflow couples liquid chromatography to mass spectrometry (LC-MS) and is centered on the high-resolution mass spectrometric analysis of HILIC-enriched and C18-LC-separated tryptic and non-tryptic intact glycopeptides. It takes advantage of the mass spectrometer’s stepped collisional energy fragmentation capabilities (HCD) and enables the unambiguous identification of the glycan moiety and the peptide moiety alike. We have identified and systematically evaluated the occurrence of a conserved N-glycopeptide fragmentation signature (Figure 2). This signature allows to unambiguously identify the peptide moiety of N-glycopeptides thus significantly increasing the reliability of the identification. Moreover, we have systematically evaluated the occurrence of unique and glycan-related fragment ion pattern (oxonium ions) that provide additional structural information on the N- or O-glycan moiety of the glycopeptides.

![Figure 2: Top Left: Glycoproteomic workflow developed for the in-depth analysis of N- and O-glycosylated proteins. Top Right: Conserved N-glycopeptide fragmentation pattern consistently obtained when using stepped normalized collisional energy (HCD.step). Bottom: HCD fragment ion spectra (MS2) of tryptic IgG 1 Fc N-glycopeptide 176EEQYNSTYR184 + HexNAc4Hex3Fuc1 (m/z 878.687 [M+3H]3+) acquired by nano-RP-LC-ESI-OT-OT-MS/MS using varying normalized collisional energies (NCE; positive ion mode, LTQ Orbitrap Elite hybrid mass spectrometer). Taken from Hoffmann et al. (2018), Proteomics, in press. Figure 2: Top Left: Glycoproteomic workflow developed for the in-depth analysis of N- and O-glycosylated proteins. Top Right: Conserved N-glycopeptide fragmentation pattern consistently obtained when using stepped normalized collisional energy (HCD.step). Bottom: HCD fragment ion spectra (MS2) of tryptic IgG 1 Fc N-glycopeptide 176EEQYNSTYR184 + HexNAc4Hex3Fuc1 (m/z 878.687 [M+3H]3+) acquired by nano-RP-LC-ESI-OT-OT-MS/MS using varying normalized collisional energies (NCE; positive ion mode, LTQ Orbitrap Elite hybrid mass spectrometer). Taken from Hoffmann et al. (2018), Proteomics, in press.](/3629800/original-1547802835.jpg?t=eyJ3aWR0aCI6MzQxLCJmaWxlX2V4dGVuc2lvbiI6ImpwZyIsIm9ial9pZCI6MzYyOTgwMH0%3D--f6d55a1aa063bfa04df2ab1b4af5dcc125e1ca66)