Generation and Mode of Action of IAV DIPs
Motivation
Influenza A virus (IAV) infections cause high morbidity and mortality in humans and animals. In principle, the error-prone RNA-dependent RNA polymerase of IAV introduces mutations during replication of the genome, facilitating virus evolution. Sporadically, large internal deletions in one of its eight viral RNA (vRNA) genome segments can occur, resulting in formation of defective interfering (DI) particles (DIPs), which are replication-defective. However, a standard virus (STV) co-infection with a DIP in a host cell leads to replication and packaging of the DI genome (Figure 1). Here, it is hypothesized that the smaller size of the DI genome allows for a faster and stronger accumulation in host cells compared to full-length counterparts. Moreover, “snatching of resources” for replication and increasing release of DIPs may lead to suppression of STV propagation. But the exact mechanisms of interference are still not fully understood. It was speculated that yet unidentified properties of DI genomes may potentially further contribute to their interfering capacity. Previous animal experiments revealed increased survival upon treatment with DI virus followed by infection with IAV. Furthermore, non-specific protection against other respiratory viruses was shown, probably mediated by the stimulation of innate immunity. Thus, DIPs have the potential to be utilized for antiviral therapy. However, in-depth molecular characterization of the interfering mechanisms is needed to develop strong candidate DIPs against virus infections.
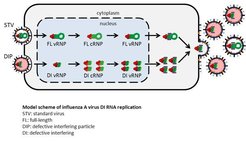
Figure 1: Scheme of DIP propagation. In a co-infection with a DIP and STV, an increased replication and preferential packaging of the DI genome can be observed, and mainly non-infectious DIPs are released. Abbreviations: FL, full length; DI defective interfering; vRNP, viral ribonucleoprotein; cRNP, complementary RNA containing ribonucleoprotein.
Aim of the project
Using reverse genetics for IAV, we aim to reconstitute “pure” candidate DIPs, identified in previous screening experiments, for detailed characterization of inference mechanisms. Therefore, we employ techniques like real-time RT-qPCR, imaging flow cytometry, and in vitro interference assays. With an improved understanding of the molecular mechanisms of DIPs, the ultimate goal is to find superior candidate DIPs or alternatively, approaches to introduce the interfering mechanisms of DIPs into cells, for the development of a prophylactic and/or therapeutic antiviral agent against influenza infections.
